
DNA polymerase
Encyclopedia
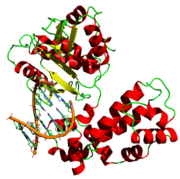
Polymerase
A polymerase is an enzyme whose central function is associated with polymers of nucleic acids such as RNA and DNA.The primary function of a polymerase is the polymerization of new DNA or RNA against an existing DNA or RNA template in the processes of replication and transcription...
is an enzyme
Enzyme
Enzymes are proteins that catalyze chemical reactions. In enzymatic reactions, the molecules at the beginning of the process, called substrates, are converted into different molecules, called products. Almost all chemical reactions in a biological cell need enzymes in order to occur at rates...
that helps catalyze in the polymerization
Polymerization
In polymer chemistry, polymerization is a process of reacting monomer molecules together in a chemical reaction to form three-dimensional networks or polymer chains...
of deoxyribonucleotide
Deoxyribonucleotide
A deoxyribonucleotide is the monomer, or single unit, of DNA, or deoxyribonucleic acid. Each deoxyribonucleotide comprises three parts: a nitrogenous base, a deoxyribose sugar, and one phosphate group. The nitrogenous base is always bonded to the 1' carbon of the deoxyribose, which is distinguished...
s into a DNA
DNA
Deoxyribonucleic acid is a nucleic acid that contains the genetic instructions used in the development and functioning of all known living organisms . The DNA segments that carry this genetic information are called genes, but other DNA sequences have structural purposes, or are involved in...
strand. DNA polymerases are best known for their feedback role
Negative feedback
Negative feedback occurs when the output of a system acts to oppose changes to the input of the system, with the result that the changes are attenuated. If the overall feedback of the system is negative, then the system will tend to be stable.- Overview :...
in DNA replication
DNA replication
DNA replication is a biological process that occurs in all living organisms and copies their DNA; it is the basis for biological inheritance. The process starts with one double-stranded DNA molecule and produces two identical copies of the molecule...
, in which the polymerase "reads" an intact DNA strand as a template and uses it to synthesize the new strand. This process copies a piece of DNA. The newly polymerized molecule is complementary to the template strand and identical to the template's original partner strand. DNA polymerases use magnesium
Magnesium
Magnesium is a chemical element with the symbol Mg, atomic number 12, and common oxidation number +2. It is an alkaline earth metal and the eighth most abundant element in the Earth's crust and ninth in the known universe as a whole...
ions as cofactors
Cofactor (biochemistry)
A cofactor is a non-protein chemical compound that is bound to a protein and is required for the protein's biological activity. These proteins are commonly enzymes, and cofactors can be considered "helper molecules" that assist in biochemical transformations....
. Human DNA polymerases are 900-1000 amino acids long.
Function
Hydroxide
Hydroxide is a diatomic anion with chemical formula OH−. It consists of an oxygen and a hydrogen atom held together by a covalent bond, and carrying a negative electric charge. It is an important but usually minor constituent of water. It functions as a base, as a ligand, a nucleophile, and a...
, and, therefore, needs a primer
Primer (molecular biology)
A primer is a strand of nucleic acid that serves as a starting point for DNA synthesis. They are required for DNA replication because the enzymes that catalyze this process, DNA polymerases, can only add new nucleotides to an existing strand of DNA...
at which it can add the first nucleotide. Primers consist of RNA
RNA
Ribonucleic acid , or RNA, is one of the three major macromolecules that are essential for all known forms of life....
and/or DNA bases. In DNA replication, the first two bases are always RNA, and are synthesized by another enzyme called primase
Primase
DNA primase is an enzyme involved in the replication of DNA.Primase catalyzes the synthesis of a short RNA segment called a primer complementary to a ssDNA template...
. An enzyme known as a helicase
Helicase
Helicases are a class of enzymes vital to all living organisms. They are motor proteins that move directionally along a nucleic acid phosphodiester backbone, separating two annealed nucleic acid strands using energy derived from ATP hydrolysis.-Function:Many cellular processes Helicases are a...
is required to unwind DNA from a double-strand structure to a single-strand structure to facilitate replication of each strand consistent with the semiconservative model of DNA replication.
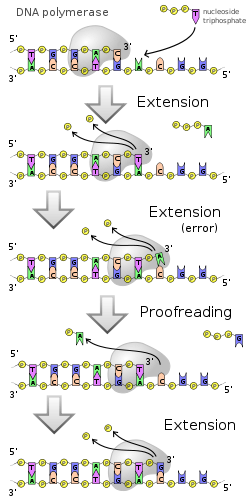
Error correction is a property of some, but not all, DNA polymerases. This process corrects mistakes in newly synthesized DNA. When an incorrect base pair is recognized, DNA polymerase reverses its direction by one base pair of DNA. The 3'-5' exonuclease
Exonuclease
Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3’ or the 5’ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle ...
activity of the enzyme allows the incorrect base pair to be excised (this activity is known as proofreading
Proofreading (biology)
The term proofreading is used in genetics to refer to the error-correcting processes, first proposed by John Hopfield and Jacques Ninio, involved in DNA replication, immune system specificity, enzyme-substrate recognition among many other processes that require enhanced specificity...
). Following base excision, the polymerase can re-insert the correct base and replication can continue.
Various DNA polymerases are extensively used in molecular biology experiments.
Variation across species
DNA polymerases have highly conserved structure, which means that their overall catalytic subunits vary, on a whole, very little from species to species. Conserved structures usually indicate important, irreplaceable functions of the cell, the maintenance of which provides evolutionary advantages.Some viruses also encode special DNA polymerases, such as Hepatitis B virus DNA polymerase
Hepatitis B virus DNA polymerase
Hepatitis B virus DNA polymerase is a hepatitis B viral protein. It encodes a DNA polymerase....
. These may selectively replicate viral DNA through a variety of mechanisms. Retroviruses encode an unusual DNA polymerase called reverse transcriptase
Reverse transcriptase
In the fields of molecular biology and biochemistry, a reverse transcriptase, also known as RNA-dependent DNA polymerase, is a DNA polymerase enzyme that transcribes single-stranded RNA into single-stranded DNA. It also helps in the formation of a double helix DNA once the RNA has been reverse...
, which is an RNA-dependent DNA polymerase (RdDp). It polymerizes DNA from a template of RNA
RNA
Ribonucleic acid , or RNA, is one of the three major macromolecules that are essential for all known forms of life....
.
DNA polymerase families
Based on sequence homology, DNA polymerases can be further subdivided into seven different families: A, B, C, D, X, Y, and RT.Family A
Polymerases contain both replicative and repair polymerases. Replicative members from this family include the extensively studied T7 DNA polymeraseT7 DNA polymerase
The T7 DNA polymerase of the T7 bacteriophage is a DNA-dependent DNA polymerase responsible for the fast rate of T7 phage DNA replication in vivo. The polymerase consists of a 1:1 complex of the viral T7 gene 5 protein and the E...
, as well as the eukaryotic mitochondrial DNA Polymerase γ. Among the repair polymerases are Escherichia coli DNA pol I, Thermus aquaticus
Thermus aquaticus
Thermus aquaticus is a species of bacterium that can tolerate high temperatures, one of several thermophilic bacteria that belong to the Deinococcus-Thermus group...
pol I, and Bacillus stearothermophilus
Bacillus stearothermophilus
Bacillus stearothermophilus is a rod-shaped, Gram-positive bacterium and a member of the division Firmicutes. The bacteria is a thermophile and is widely distributed in soil, hot springs, ocean sediment, and is a cause of spoilage in food products. It will grow within a temperature range of 30-75...
pol I. These repair polymerases are involved in excision repair and processing of Okazaki fragment
Okazaki fragment
Okazaki fragments are short molecules of single-stranded DNA that are formed on the lagging strand during DNA replication. They are between 1,000 to 2,000 nucleotides long in Escherichia coli and are between 100 to 200 nucleotides long in eukaryotes....
s generated during lagging strand synthesis.
Family B
B family polymerases are involved in DNA repair and replication. These include; Pol II (bacterial), Pol B (archaebacterial), and Pol α, δ, and ε (eukaryotic). Pol ζ another B family polymerase, is made of two subunits Rev3 and Rev7 and is involved in translesion synthesis. Pol ζ is unique in that it can extend primers with terminal mismatches.Family C
Polymerases are the primary bacterial chromosomal replicative enzymes. DNA Polymerase III alpha subunit from E. coli is the catalytic subunit and possesses no known nuclease activity. A separate subunit, the epsilon subunit, possesses the 3'-5' exonuclease activity used for editing during chromosomal replication. Recent research has classified Family C polymerases as a subcategory of Family X.Family D
Polymerases are still not very well characterized. All known examples are found in the EuryarchaeotaEuryarchaeota
In the taxonomy of microorganisms, the Euryarchaeota are a phylum of the Archaea.The Euryarchaeota include the methanogens, which produce methane and are often found in intestines, the halobacteria, which survive extreme concentrations of salt, and some extremely thermophilic aerobes and anaerobes...
subdomain of Archaea and are thought to be replicative polymerases.
Family X
Contains the well-known eukaryotic polymerase pol β, as well as other eukaryotic polymerases such as pol σ, pol λ, pol μ, and terminal deoxynucleotidyl transferase (TdT). Pol β is required for short-patch base excision repairBase excision repair
In biochemistry and genetics, base excision repair is a cellular mechanism that repairs damaged DNA throughout the cell cycle. It is responsible primarily for removing small, non-helix-distorting base lesions from the genome. The related nucleotide excision repair pathway repairs bulky...
, a DNA repair pathway that is essential for repairing alkylated or oxidised bases as well as abasic sites. Pol λ and Pol μ are involved in non-homologous end-joining, a mechanism for rejoining DNA double-strand breaks. TdT is expressed only in lymphoid tissue, and adds "n nucleotides" to double-strand breaks formed during V(D)J recombination
V(D)J recombination
VJ recombination, also known as somatic recombination, is a mechanism of genetic recombination in the early stages of immunoglobulin and T cell receptors production of the immune system...
to promote immunological diversity. The yeast Saccharomyces cerevisiae
Saccharomyces cerevisiae
Saccharomyces cerevisiae is a species of yeast. It is perhaps the most useful yeast, having been instrumental to baking and brewing since ancient times. It is believed that it was originally isolated from the skin of grapes...
has only one Pol X polymerase, Pol IV, which is involved in non-homologous end-joining.
Family Y
Y Polymerases differ from others in having a low fidelity on undamaged templates and in their ability to replicate through damaged DNA, known as translesion synthesis (TLS). Members of this family are hence called translesion synthesis polymerases. Depending on the lesion, TLS polymerases can bypass the damage in an error-free or error-prone fashion, the latter resulting in elevated mutagenesis. Xeroderma pigmentosumXeroderma pigmentosum
Xeroderma pigmentosum, or XP, is an autosomal recessive genetic disorder of DNA repair in which the ability to repair damage caused by ultraviolet light is deficient. In extreme cases, all exposure to sunlight must be forbidden, no matter how small. Multiple basal cell carcinomas and other skin...
variant (XPV) patients for instance have mutations in the gene encoding Pol η (eta)
DNA polymerase eta
DNA polymerase eta is a eukaryotic DNA polymerase involved in the DNA repair by translesion synthesis. The gene encoding DNA polymerase eta is POLH, also known as XPV, because loss of this gene results in the disease Xeroderma Pigmentosum Variant...
, which is error-free for UV-lesions. In XPV patients, alternative error-prone polymerases, e.g., Pol ζ (zeta) (polymerase ζ is a B Family polymerase a complex of the catalytic subunit REV3L
REV3L
Protein reversionless 3-like also known as DNA polymerase zeta catalytic subunit is an enzyme that in humans is encoded by the REV3L gene....
with Rev7, which associates with Rev1
REV1
DNA repair protein REV1 is a protein that in humans is encoded by the REV1 gene. Rev1 is a Y family DNA polymerase; it is sometimes referred to as a deoxycytidyl transferase because it only inserts deoxycytidine across from lesions. Whether G, A, T, C, or an abasic site, Rev1 will always add a C...
), are thought to be involved in mistakes that result in the cancer predisposition of these patients. Other members in humans are Pol ι (iota)
POLI
DNA polymerase iota is an enzyme that in humans is encoded by the POLI gene. It is found in higher eukaryotes, and is believed to have arisen from a gene duplication from Pol η. Pol ι, is a Y family polymerase that is involved in translesion synthesis. It can bypass 6-4 pyrimidine adducts and...
, Pol κ (kappa), and Rev1
REV1
DNA repair protein REV1 is a protein that in humans is encoded by the REV1 gene. Rev1 is a Y family DNA polymerase; it is sometimes referred to as a deoxycytidyl transferase because it only inserts deoxycytidine across from lesions. Whether G, A, T, C, or an abasic site, Rev1 will always add a C...
(terminal deoxycytidyl transferase). In E. coli, two TLS polymerases, Pol IV (DINB) and Pol V (UmuD'2C), are known.
Family RT
The reverse transcriptase family contains examples from both retroviruses and eukaryotic polymerases. The eukaryotic polymerases are usually restricted to telomeraseTelomerase
Telomerase is an enzyme that adds DNA sequence repeats to the 3' end of DNA strands in the telomere regions, which are found at the ends of eukaryotic chromosomes. This region of repeated nucleotide called telomeres contains non-coding DNA material and prevents constant loss of important DNA from...
s. These polymerases use an RNA template to synthesize the DNA strand.
Prokaryotic DNA polymerases
Bacteria have 5 known DNA polymerases.Eukaryotic DNA polymerases
Eukaryotes have at least 15 DNA Polymerases.External links
- Unusual repair mechanism in DNA polymerase lambda, Ohio State UniversityOhio State UniversityThe Ohio State University, commonly referred to as Ohio State, is a public research university located in Columbus, Ohio. It was originally founded in 1870 as a land-grant university and is currently the third largest university campus in the United States...
, July 25, 2006. - A great animation of DNA Polymerase from WEHI at 1:45 minutes in

